Gene expression is a process of using genetic information within a gene to produce a functional gene product such as RNA or protein. Microarray and next generation RNA-sequencing (RNA-seq) are two main assays used in the study of gene expression profile at transcriptome level. A recent post by GeneVia technologies suggests a quick search on google trends would show a steady drop in the demand for microarray application over the years with the emergence of RNA-Seq. This illustrates a technology shift from the 24 years old microarray to the more recent RNA-Seq technology. Microarray is a technique which relies on the relative intensity of fluorescently tagged complementary DNA (cDNA) to determine gene expressions while RNA-seq depends on high-throughput RNA sequencing data which enables researchers to obtain information about the transcriptome of a cell to analyze data at Basepair.
Microarray protocol in brief involves four basic steps. First, the mRNA from a particular gene of interest is isolated and purified. Then, the mRNA is reverse transcribed with the help of reverse transcriptase enzyme to produce ds-cDNA which are then fluorescently tagged. These tagged cDNA hybridizes to complementary, pre-designed synthetic oligonucleotides immobilized onto microscopic spots within a solid surface called DNA chip. The fourth step involves scanning of the microarray. The intensity of the fluorescent across the array indicates the level of gene expression of a particular gene within the genome of interest. On the other hand, in RNA-Seq technology, the RNA will be isolated and converted to cDNA libraries. These libraries will then be sequenced using next-generation sequencing platforms to produce high quality RNA sequencing data in the form of reads. These RNA sequencing data will be aligned and used to determine gene expressions of the gene of interest.
Hybridization-based microarray technology has been a robust method widely used in the study of gene expression over the years while studies using RNA sequencing data is an advancing technology in the more recent years. Comparing these two methods will give us a clearer idea of which method is more advantageous and reliable for gene expression studies. To begin with, the key consideration in every study is the cost of research itself. Hence, it is important to strike a balance between the cost and performance of the chosen method. Microarray has been favoured for its advantage of being a high-throughput yet relatively low cost technology. Despite the drop in sequencing prices, generating next generation RNA sequencing data is still more expensive compared to microarray. This makes microarray significantly more economical when it comes to working on large projects with big number of samples.
However, looking into other key differences would easily outweigh the cost factor of RNA-seq. RNA sequencing data allows for the study of non-model organisms, novel genes and structural variants. In contrast, microarray is not suitable for the study of non-model organisms as it requires prior sequence knowledge. A DNA chip needs to be developed for the particular genome of interest in the case of microarray. RNA-seq on the other hand gives a comprehensive view of a transcriptome. It works well for both novel and known transcripts and is suitable for discovery-based researches whereby structural variants can be identified. When dealing with new discovery, sequencing data can be reanalysed whereas microarray technology requires rerunning the procedure to understand any new discoveries.
In addition, microarray has reduced sensitivity due to its hybridization-based technology. It cannot detect differences between very similar sequences, for instance, isoforms. Microarray gives raise to problematic cross-hybridization when dealing with highly similar sequences (Kukurba and Montgomery, 2015). It also has limited ability to accurately quantify high and low gene expression levels. Microarray measures probe intensity to determine levels of gene expression while RNA-seq quantifies aligned digital read counts. Since each and every transcript is being sequenced, RNA-seq is more sensitive in detecting low levels of expressions and isoforms.
Next, RNA-seq has lower noise compared to microarray. This is because the sequenced transcripts can be explicitly mapped onto unique regions of a genome enabling background signals to be eliminated easily during analysis. Besides, errors due to cross-hybridization can also be removed through RNA-seq method. Another advantage of RNA-seq over microarray is that it can accommodate comparison of large number of samples and not just limited to fluorescent tagging of individual samples. In terms of data analysis, though sequencing generates much richer information which might result in a more complex analysis, there are many user-friendly tools being continuously developed to help ease RNA-seq data analysis and interpretation even for researchers without bioinformatics background.
Due to the disadvantages of microarray, there may be circumstances where one may start out with microarray and eventually end up using RNA-seq. This will only result in extra expenses. On the whole, even though microarray is a more economical technology, RNA-seq will be a better and more reliable choice for gene studies due to its aforementioned advantages over microarray technology.
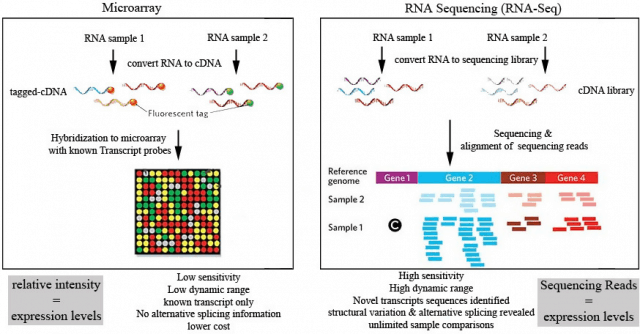
Source of image: otogenetics.com
This post has been sponsored by NearFox
Digital Health Buzz!
Digital Health Buzz! aims to be the destination of choice when it comes to what’s happening in the digital health world. We are not about news and views, but informative articles and thoughts to apply in your business.


